In this continuing series, we explore the NMath Stats functions for performing cluster analysis. (For previous posts, see Part 1 – PCA and Part 2 – K-Means.) The sample data set we’re using classifies 89 single malt scotch whiskies on a five-point scale (0-4) for 12 flavor characteristics. To visualize the data set and clusterings, we make use of the free Microsoft Chart Controls for .NET, which provide a basic set of charts.
In this post, we’ll cluster the scotches based on “similarity” in the original 12-dimensional flavor space using hierarchical cluster analysis. In hierarchical cluster analysis, each object is initially assigned to its own singleton cluster. The analysis then proceeds iteratively, at each stage joining the two most “similar” clusters into a new cluster, continuing until there is one overall cluster. In NMath Stats, class ClusterAnalysis performs hierarchical cluster analyses.
The clustering process is governed by two functions:
- A distance function computes the distance between individual objects. In NMath Stats, the distance function is encapsulated in a Distance.Function delegate, which takes two vectors and returns a measure of the distance (similarity) between them. Delegates are provided as static variables on class Distance for many common distance functions. You can also define your own delegate.
- A linkage function computes the distance between clusters. In NMath Stats, the linkage function is encapsulated in a Linkage.Function delegate. When two groups P and Q are united, a linkage function computes the distance between the new combined group P + Q and another group R. Delegates are provided as static variables on class Linkage for many common linkage functions. Again, you can also define your own delegate.
Based on the choice of distance and linkage function, radically different clustering can often result. Ultimately, background knowledge of the domain is required to choose between them.
In this case, we’ll use the Euclidean distance function and the Ward linkage function. The Ward linkage function computes the distance between two clusters using Ward’s method, which tends to produce compact groups of well-distributed size. Ward’s method uses an analysis of variance approach to evaluate the distances between clusters. The smaller the increase in the total within-group sum of squares as a result of joining two clusters, the closer they are. The within-group sum of squares of a cluster is defined as the sum of the squares of the distance between all objects in the cluster and the centroid of the cluster.
This code clusters the scotch data (loaded into a DataFrame in Part I), then cuts the hierarchical cluster tree at the level of four clusters:
ClusterAnalysis ca = new ClusterAnalysis(df,
Distance.EuclideanDistance,
Linkage.WardFunction
);
ClusterSet cs = ca.CutTree(4);
Printing out the cluster members from the cluster set, as shown in Part II, produces:
Objects in cluster 0: Aberfeldy Aberlour Ardmore Auchroisk Balmenach Belvenie BenNevis Benriach Benrinnes Benromach BlairAthol Dailuaine Dalmore Deanston Edradour GlenElgin GlenKeith GlenOrd Glendronach Glendullan Glenfarclas Glenlivet Glenrothes Glenturret Knochando Linkwood Longmorn Macallan Mortlach OldFettercairn RoyalBrackla RoyalLochnagar Strathisla Tullibardine Objects in cluster 1: AnCnoc ArranIsleOf Auchentoshan Aultmore Bladnoch Bunnahabhain Cardhu Craigallechie Dalwhinnie Dufftown GlenDeveronMacduff GlenGrant GlenMoray GlenSpey Glenallachie Glenfiddich Glengoyne Glenkinchie Glenlossie Glenmorangie Inchgower Loch Lomond Mannochmore Miltonduff Scapa Speyburn Speyside Tamdhu Tobermory Tomintoul Tomore Objects in cluster 2: Ardbeg Caol Ila Clynelish Lagavulin Laphroig Talisker Objects in cluster 3: Balblair Bowmore Bruichladdich Craigganmore GlenGarioch GlenScotia Highland Park Isle of Jura Oban OldPulteney Springbank Strathmill Tamnavulin Teaninich Tomatin
Coloring the objects based on cluster assignment in the plot of the first two principal components shows how similar this clustering is to the results of k-mean clustering.
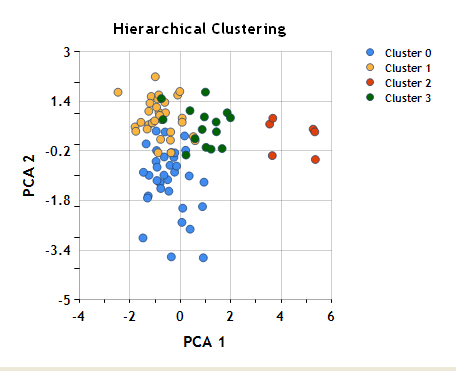
Again, remember that although we’ve used dimension reduction (principal component analysis, in this case) to visualize the clustering, the clustering itself was performed based on similarity in the original 12-dimensional flavor space, not based on distance in this plane.
Because we have the entire hierarchical cluster tree, we can cut the tree at different levels. For example, into six clusters:
Objects in cluster 0: Aberfeldy Aberlour Ardmore Auchroisk Belvenie BenNevis Benriach Benrinnes Benromach BlairAthol Deanston Edradour GlenElgin GlenKeith GlenOrd Glendullan Glenfarclas Glenlivet Glenrothes Glenturret Knochando Linkwood Longmorn OldFettercairn RoyalBrackla Strathisla Tullibardine Objects in cluster 1: AnCnoc ArranIsleOf Auchentoshan Aultmore Bladnoch Bunnahabhain Cardhu Craigallechie Dalwhinnie Dufftown GlenDeveronMacduff GlenGrant GlenMoray GlenSpey Glenallachie Glenfiddich Glengoyne Glenkinchie Glenlossie Glenmorangie Inchgower Loch Lomond Mannochmore Miltonduff Scapa Speyburn Speyside Tamdhu Tobermory Tomintoul Tomore Objects in cluster 2: Ardbeg Caol Ila Clynelish Lagavulin Laphroig Talisker Objects in cluster 3: Balblair Craigganmore GlenGarioch Oban Strathmill Tamnavulin Teaninich Objects in cluster 4: Balmenach Dailuaine Dalmore Glendronach Macallan Mortlach RoyalLochnagar Objects in cluster 5: Bowmore Bruichladdich GlenScotia Highland Park Isle of Jura OldPulteney Springbank Tomatin
Coloring the objects based on cluster assignment in the plot of the first two principal components:
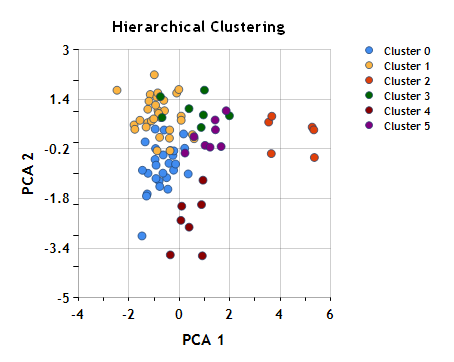
Clusters (1, 2) are unchanged, while clusters (0, 3) are now split into sub-clusters.
Both of the clustering techniques we’ve looked at so far–k-means and hierarchical cluster analysis–have clustered the scotches based on similarity in the original 12-dimensional flavor space. In the next post, we look at clustering using non-negative matrix factorization (NMF), which can be used to cluster the objects using a reduced set of synthetic dimensions.
Ken

I was hoping to see a dendgram + heatmap for your HCA results. Any plans?
Hi Johann,
I would have liked to show a dendrogram and heat map, but I don’t think there is support for these chart types in the free MS Chart package. Of course, the NMath ClusterAnalysis object contains all the information you would need to draw such charts.
More full-featured commercial charting packages usually have these chart types. For example, NMath works well with Infragistics (http://www.infragistics.com/), Syncfusion (http://www.syncfusion.com/), Nevron (http://www.nevron.com/), and SoftwareFX (http://www.softwarefx.com/), to name a few.
In the future, we will have a blog entry showing a heat map and other examples of charting using NMath.
Ken
As a follow up,
we wrote a routine to compute the coordinates needed to draw a dendrogram for hierarchical clustering results to help out Johann. It’s more complicated than one might expect at first blush.
If any other folks need it, just send us a request and we’d be happy to provide it. As the code is specifically for drawing, it probably won’t be released with the NMath Stats product.
Paul
I was wondering which of the clustering methods of the 4 or more included in NMath is most appropriate to use with large datasets, e.g. 1 million points with expected 10-20 clusters?